#12,531
Were it not for a record setting 5th wave of H7N9 cases in China, and Europe seeing their worst avian epizootic (HPAI H5N8) in history - H5N6, which first emerged in 2014 - would have garnered a lot more attention than it has over the past 6 months.
While the number of known human H5N6 infections (n=17) pales compared to H7N9 (1500+), more than half of those have proved fatal. And like H7N9, the H5N6 virus is quite promiscuous - reassorting with other avian viruses - reinventing itself as it spreads.
Influenza A viruses are broadly categorized by two proteins they carry on their surface; their HA (hemagglutinin) and NA (neuraminidase), producing subtypes like H5N1, H7N9, or H5N6. There are currently 18 known HA subtypes and 11 known NA subtypes.
When we talk about a subtype, however, we're aren't just talking about a single viral threat, but rather an expanding array of related viruses sharing the same (or similar) HA and NA genes.Within each HA subtype there are genetic groupings called clades, and often within each clade - subclades. Within each of these, many minor variants may exist. Add in the ability to mix-and-match internal genes, and you can come up with literally dozens of genotypes for each subtype.
Even minor differences within an influenza subtype can alter a virus's behavior, including infectivity and pathogenicity (see Differences In Virulence Between Closely Related H5N1 Strains). This may help explain why South Korea - despite seeing hundreds of H5N6 poultry outbreaks this past winter - reported no human cases.
Last December, in Cell Host Microbe: Genesis, Evolution and Prevalence of HPAI H5N6 In China, we saw a report that found that H5N6 had become the dominant HPAI H5 virus in Chinese ducks (replacing H5N1), with 34 distinct H5N6 genotypes, including 4 that have infected people.
While we've not seen a reported human infection with H5N6 since two cases were announced last November (link), over the winter the virus spread from China to South Korea (where it reassorted into several new genotypes), Japan, and Taiwan (see Korea H5N6: New Genetic Analysis & Investigating Its Rapid Spread).Last week, the journal Emerging Microbes & Infections published a review of the last two human cases in China (in Guangxi & Hunan provinces), which identifies a novel reassorted H5N6 behind both cases, illustrating the continual evolution of the H5N6 subtype.
I've only included the highlights, follow the link to read the full, open-access report.
Human infections with novel reassortant H5N6 avian influenza viruses in China(Continue . . . )
Open
Ye Zhang1,*, Minmei Chen2,*, Yiwei Huang3,*, Wenfei Zhu1,*, Lei Yang1, Lidong Gao3, Xiaodan Li1, Fuyin Bi2, Chaoyang Huang3, Ning Kang2, Hengjiao Zhang3, Zi Li1, Hong Bo1, Dayan Wang1 and Yuelong Shu1
Received 4 February 2017; Revised 23 March 2017; Accepted 28 March 2017
Dear Editor,
In May 2014, China officially reported the first human infection with highly pathogenic H5N6 subtype avian influenza virus (AIV) in Sichuan Province.1 Through 2nd Feb, 17 H5N6 cases were documented (http://www.who.int/en/), and H5N6 viruses isolated from both humans and poultry showed marked reassortment.2, 3, 4 We have previously identified three major reassortant H5N6 viruses, including reassortants A, B and C.4 The viruses with these three reassortant forms have caused human infections.
In November 2016, Hunan and Guangxi provinces reported two additional H5N6 human cases, one from each province. Both human cases were characterized by severe pneumonia and resulted in fatal outcomes. Epidemiologic investigations revealed that both patients had a history of exposure to dead chickens. The Chinese National Influenza Center isolated H5N6 viruses in a biosafety level 3 facility by inoculating 0.2 mL of each original sample into 9- to 11-day-old specific pathogen-free (SPF) embryonated chicken egg allantoic cavities. The isolated viruses were termed A/Hunan/55555/2016 (H5N6, HN555) and A/Guangxi/55726/2016 (H5N6, GX726).
(SNIP)
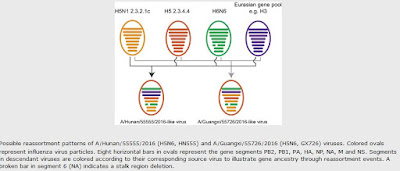
In summary, two additional human infections with highly pathogenic influenza A (H5N6) viruses were reported in November 2016 in mainland China. Both homology analysis of nucleotide sequences and phylogenetic analysis of two isolated H5N6 viruses showed that the HN555 virus represented a novel H5N6 genotype and may have been generated by the reassortment of circulating H5N6 viruses with other low-pathogenicity H3 viruses (Figure 1).
GX726 represented another novel H5N6 genotype member that has been transmitted from avian species to humans. Our findings suggest that the currently circulating H5N6 viruses in birds may continuously reassort with other enzootic low-pathogenicity influenza viruses. These will likely not be the last novel H5N6 genotype viruses to be generated. Therefore, enhanced surveillance of influenza viruses in birds and humans is continuously needed.
Up until 2013, we really only had one serious avian flu threat in the wild; H5N1. Since then, we've seen a steady procession of new contenders emerge - mostly from China (H7N9, H5N8, H5N6, H10N8, etc.).
H7N9 currently sits atop the world's pandemic watch list (see NPR: A Pessimistic Guan Yi On H7N9's Evolution), but the fortunes of flu viruses can change very quickly. Just a year ago, H7N9 appeared to be on the decline, posting its lowest number of human infections since it emerged in 2013.
One year later, after a new (Yangtze River Delta) lineage and an HPAI version emerged, it's suddenly a new ball game.Despite the warning signs, we've an astonishingly bad record of reading the tea leaves and predicting the next big public health threat It may very well be H7N9, but the speed of viral evolution (particularly via reassortment) is such that we can easily be blindsided by a threat we never saw coming.
The authors of today's report end by warning; `These will likely not be the last novel H5N6 genotype viruses to be generated'.Of course, the same can be said for practically every other novel flu subtype in the wild. Which is why pandemic preparedness needs to be not only a national priority, but a global one as well (see World Bank: World Ill-Prepared For A Pandemic).