How viruses shuffle their genes (reassort)
# 9912
After more than 17 years of relative stability (1996-2013) - during which time we only had one major HPAI H5 avian virus (H5N1) to concern ourselves with - we’ve seen a sudden and unprecedented expansion of highly pathogenic H5 avian subtypes around the globe.
The first minor crack in H5’s veneer appeared in 2008, when two mallard ducks in Eastern China tested positive for a new subtype H5N5 (see Novel H5N5 Avian Influenza Detected In China).
The account of its discovery appeared in the 2011 EID Journal Dispatch called Novel Reassortant Highly Pathogenic Avian Influenza (H5N5) Viruses in Domestic Ducks, China, where they identified the likely parental viruses (H5N1 and H6N5) both circulating in local domestic ducks.
While never a huge `player’ in the avian flu world, H5N5 demonstrated that H5N1 could reassort into a novel subtype, and suggested that domestic ducks could serve as `mixing’ vessels for creating new subtypes of influenza viruses.
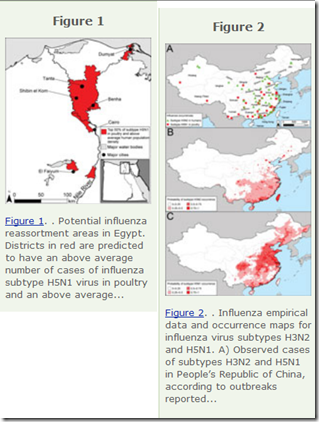
In an instance of particularly good timing – in the middle of March of 2013, just two weeks before we learned of the emergence of H7N9 in China – the EID Journal published a research Article on Predicting Hotspots for Influenza Virus Reassortment.
While the northern plains of India, the western Korean Peninsula and southwestern Japan were mentioned, their two biggest hotspots were Eastern China, and the Nile Valley of Egypt – both regions that have produced either new subtypes, or new clades (see Emergence Of A Novel Cluster of H5N1 Clade 2.2.1.2), of HPAI H5 viruses over the past two years.
In January of 2014, an emerging HPAI H5N8 appeared in South Korea, and rapidly spread through that nation’s poultry and wild bird population. It showed up in Japan in April, and was subsequently reported in China.
During the winter of 2014-15, H5N8 migrated to Russia, Western Europe, and even North America – and along the way spawned additional reassortants (H5N2, H5N3, H5N1) when it mixed with local avian flu subtypes. Thus far, none of these H5N8 derived viruses have proven pathogenic in humans.
During the Spring of 2014 another HPAI H5 appeared in Southeast Asia; H5N6. Unlike the H5N8 virus (and its descendents), H5N6 has caused serious (even fatal) human illness (see China Reports 3rd H5N6 Case (Fatal) – Yunnan Province).
Suddenly we’ve gone from one HPAI H5 virus of concern, to a half dozen. And as these viruses spread, and mingle with other viruses, more novel subtypes may yet emerge. And as the following dispatch from the EID Journal points out, their future behavior may be unpredictable.
Dispatch
Erik de Vries
, Hongbo Guo1, Meiling Dai1, Peter J.M. Rottier, Frank J.M. van Kuppeveld, and Cornelis A.M. de Haan Abstract
In 2014, novel highly pathogenic avian influenza A H5N2, H5N5, H5N6, and H5N8 viruses caused outbreaks in Asia, Europe, and North America. The H5 genes of these viruses form a monophyletic group that evolved from a clade 2.3.4 H5N1 variant. This rapid emergence of new H5Nx combinations is unprecedented in the H5N1 evolutionary history.
A highly pathogenic avian influenza (HPAI) A(H5N1) virus (A/goose/Guangdong/1/1996) was first detected in China in 1996. Multiple clades, defined by phylogenetic characterization of the H5 hemagglutinin (HA) (1), have evolved and spread across Asia, Africa, and Europe, causing enormous losses to the poultry industry. A total of 694 human infections (death rate 58%) were recorded during 2003–2014 (2).
During the evolution of HPAI H5N1 viruses, reassortment events involving the 6 internal gene segments have often been detected (reviewed in [3]), but novel subtypes (i.e., combinations of HPAI H5 with other N subtypes) have rarely been isolated. In 2014, a novel highly virulent reassortant HPAI H5N6 virus (4) caused multiple outbreaks in Southeast Asia and 1 lethal human infection, which led the Food and Agricultural Organization of the United Nations to issue a warning (5). Outbreaks of novel HPAI H5N8 virus in South Korea (6,7), China (8), and Japan raised further concern, and in November 2014, this subtype emerged outside Eastern Asia, causing outbreaks in poultry farms in Germany, the Netherlands, the United Kingdom, Canada, and the United States.
<SNIP>
Conclusion
(Excerpt)
In this study, we exclusively focused on the unique occurrence of new HA–NA combinations. Recent publications have already described the reassortment events of the internal gene segments of several of the viruses mentioned above (6–8,11–14). In contrast to novel HA–NA combinations, novel constellations of internal gene segments are far from unique and have frequently been observed for HPAI H5N1 viruses (3). Our analysis indicates that new HPAI viruses have emerged that carry H5 proteins capable of matching with multiple NA subtypes. Whether the formation of new HA–NA combinations confers a selective advantage that contributed to the emergence of these novel subtypes is not known and requires elaborate research. However, the balance between HA (receptor binding) and NA (receptor cleavage) protein activities is known to be critical to cell entry and host tropism and may be an important factor that lead to the emergence of new HA–NA combinations. In contrast to HPAI H5N1, the novel clade 2.3.4.4 viruses, excluding H5N6 viruses, have not caused human infections. However, it is unknown to what extent the repeated acquisition of a new NA proteins could enhance the rate of evolution of the HA protein. Obviously such changes could further affect host and tissue specificity, potentially having serious consequences. Therefore, surveillance is required to monitor further spread, evolution, and potential changes in host range.
Given the recent emergence of H5N8, H5N6, H5N3 in Asia and novel reassortants of H5N2 and H5N1 in North America, and it comes as little surprise that the World Health Organization recently released a pointed warning that H5 Is Currently The Most Obvious Avian Flu Threat.