#7851
In March of 2012, researchers from the CDC and the Universidad del Valle of Guatemala announced they had found the first evidence of influenza infection in bats (see A New Flu Comes Up To Bat). With the addition of an H17 flu subtype, and a new host species, suddenly all of the textbooks and slide presentations on influenza were out of date once more.
(A bit of history: the H16 subtype was discovered in 2004 by a team that included Ron Fouchier).
While preliminary research suggested that while this H17 virus was genetically compatible with human influenza viruses, it did not yet have the ability to infect humans. The CDC wrote:
As a result, it is possible that these bat viruses could eventually gain the ability to cause infections in humans. However, CDC experts have been unable to grow the bat influenza virus in test tubes, suggesting that the virus is currently not well suited to causing human illness. For the bat influenza virus to infect humans, it would need to obtain some genetic properties of human influenza viruses
The most likely scenario for this to occur would be through the reassortment of a bat and a human virus in an intermediate host, such as a pig.
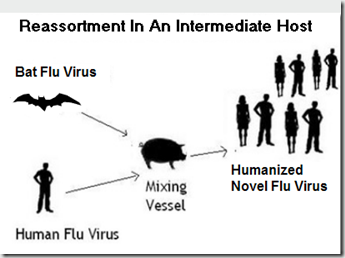
The CDC’s Bat Flu Q & A has this to say about the possibility of such a reassortment occurring:
However, the conditions needed for reassortment to occur between human influenza viruses and bat influenza virus remain unknown. A different animal (such as pigs, horses or dogs) would need to serve as a “bridge,” meaning that such an animal would need to be capable of being infected with both this new bat influenza virus and human influenza viruses for reassortment to occur. Additional studies are needed to determine the likelihood that reassortment would occur in nature between bat and human influenza viruses.
Today, via the open access journal PLoS Pathogens, we learn that different team of researchers has discovered another flu subtype – classified as H18N11 – circulating in bats sampled in Peru.
Research Article
New World Bats Harbor Diverse Influenza A Viruses
Suxiang Tong mail, Xueyong Zhu, Yan Li, Mang Shi, Jing Zhang, Melissa Bourgeois, Hua Yang, Xianfeng Chen, Sergio Recuenco, Jorge Gomez, Li-Mei Chen, Adam Johnson, Ying Tao, [ ... ], Ruben O. Donis mail
Abstract
Aquatic birds harbor diverse influenza A viruses and are a major viral reservoir in nature. The recent discovery of influenza viruses of a new H17N10 subtype in Central American fruit bats suggests that other New World species may similarly carry divergent influenza viruses. Using consensus degenerate RT-PCR, we identified a novel influenza A virus, designated as H18N11, in a flat-faced fruit bat (Artibeus planirostris) from Peru. Serologic studies with the recombinant H18 protein indicated that several Peruvian bat species were infected by this virus. Phylogenetic analyses demonstrate that, in some gene segments, New World bats harbor more influenza virus genetic diversity than all other mammalian and avian species combined, indicative of a long-standing host-virus association. Structural and functional analyses of the hemagglutinin and neuraminidase indicate that sialic acid is not a ligand for virus attachment nor a substrate for release, suggesting a unique mode of influenza A virus attachment and activation of membrane fusion for entry into host cells. Taken together, these findings indicate that bats constitute a potentially important and likely ancient reservoir for a diverse pool of influenza viruses.
The above is just small excerpt from the study, you’ll want to follow the link to read it in its entirety. Among their findings:
- A high seroprevalence of influenza antibodies in bats they tested, suggesting `widespread circulation of influenza A viruses among New World bats’.
- The author’s also found, that unlike human influenza viruses, `The crystal structures of the hemagglutinin and neuraminidase proteins indicate that sialic acid is not a receptor for virus attachment nor a substrate for release, suggesting a novel mechanism of influenza A virus attachment and activation of membrane fusion for entry into host cells.’
The authors summarize their findings in the discussion section:
Multiple lines of evidence suggest that influenza viruses have evolved in bats for an extended period of time: (i) in four of the eight gene segments, genetic diversity exceeds that observed in all other animal species combined; (ii) the divergence into multiple HA subtypes and utilization of alternative mechanisms for sialic acid-independent virion attachment to target cells and subsequent release; and (iii) the widespread geographic distribution in the Americas (Guatemala and Peru sampling sites are ~3,500 km apart) combined with a high seroprevalence in several bat species are indicative of an established infection, while the observation that the bat viruses form a monophyletic group is suggestive of sustained transmission in this species. The postulated ancient relationship of influenza viruses with aquatic migratory birds is consistent with the optimized parasitic relationship of the virus with ducks, involving subclinical infections with multiple virus subtypes as a result of efficient fecal-oral transmission. Although necessarily preliminary in nature, the data presented here suggest that similar ecological and evolutionary strategies may have been exploited by the influenza A viruses of New World bats.
All things considered, the past couple of decades have turned out to be busy ones for Chiroptologists (scientists who study bats). These winged mammals are increasingly being viewed as naturals hosts for, and potential vectors of, a number of newly recognized emerging pathogens.
Long known for vectoring rabies, in the 1990s bats gained notoriety with the emergence of the Hendra virus in Australia in 1994 (see Australia: Hendra Vaccine Hurdles) and Nipah in Malaysia in 1999 (see MMWR Update: Outbreak of Nipah Virus -- Malaysia and Singapore, 1999).
The emergence of the SARS coronavirus in 2003 – ultimately linked to bats – and most recently, to the MERS coronavirus in the Middle East (see EID Journal: Detection Of MERS-CoV In Saudi Arabian Bat), has served to cement their reputation as important carriers of emerging infectious diseases.
None of this is meant to demonize bats, as they play an important role in our ecosystem. However, bats are increasingly being associated with diseases deadly to humans, so a degree of caution is warranted.
To learn how you can stay safe around bats, the CDC offers the following advice.
