#17,339
The hottest topic in the world of pandemic flu right now involve the repeated spillover of avian H5N1 into mammals around the world, and the resultant host-adaptations that may increase the virus's threat to humans.
While still primarily a threat to avian species, thousands of terrestrial and marine mammals have been infected over the past couple of years, with many experiencing severe (and often fatal) neurological manifestations, including:
USDA Update On H5N1 In Mammalian WildlifeAvian flu viruses are normally well adapted to attack the kind of cells commonly found within the avian gastrointestinal tract; α2,3-linked sialic acid avian receptor cells and to replicate efficiently at the higher temperatures found in the avian gut.
Peru:Reports Nearly 3,500 Sea Lions Killed By H5N1 Avian Flu
HPAI A(H5N1) Virus Infection in Farmed Minks, Spain, October 2022
Ontario: CWHC Reports HPAI H5 Infection With Severe Neurological Signs In Wild Foxes (Vulpes vulpes)
Netherlands DWHC Reports another Mammal (Polecat) Infected With H5N1
In order to infect and transmit efficiently among mammals, avian viruses need to be able to attach to the α2,6-linked receptor cells commonly found in their respiratory tract, and to replicate at the lower temperatures found there.
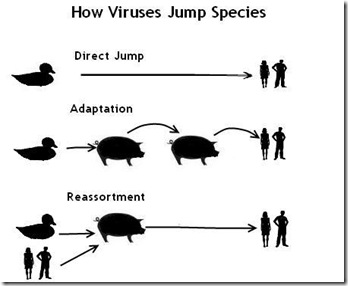
Many avian viruses have an affinity for both avian and mammalian receptor cells, which is why they are occasionally able to jump species. Once in a mammalian host, however, further `host adaptations' are needed for the virus to flourish.
- One of the key mutations that we look for is PB2-E627K; the swapping out of Glutamic acid (E) for Lysine (K) at position 627 in the PB2 protein, which allows the virus to replicate at a lower temperature.
- The PB2-D701N mutation has frequently been linked increased severity of infection (see here and here).
- There are a number of others (including T271A, K389R, D622G, and N319K), and likely many we aren't aware of their importance yet.
Of note, there were four different genotypes among the viruses sampled, illustrating the continual evolution of the virus after it arrived in North America a little over a year ago.
Due to its length I've only posted a few excerpts. You'll definitely want to follow the link to read it in its entirety.
Research Article
Tamiru N. Alkie,Sherri Cox,Carissa Embury-Hyatt,Brian Stevens,Neil Pople,Margo J. Pybus, show all
07 Mar 2023
Download citation https://doi.org/10.1080/22221751.2023.2186608Abstract
The GsGd lineage (A/goose/Guangdong/1/1996) H5N1 virus was introduced to Canada in 2021/2022 through the Atlantic and East Asia-Australasia/Pacific flyways by migratory birds. This was followed by unprecedented outbreaks affecting domestic and wild birds, with spillover into other animals.
In the current study, we report sporadic cases of H5N1 in 40 free-living mesocarnivore species such as red foxes, striped skunks and mink in Canada. The clinical presentations of the disease in mesocarnivores were consistent with central nervous system infection. This was supported by the presence of microscopic lesions and presence of abundant IAV antigen by immunohistochemistry.
Some red foxes that survived clinical infection developed anti-H5N1 antibodies. Phylogenetically, the H5N1 viruses from the mesocarnivore species belonged to clade 2.3.4.4b and had four different genome constellation patterns.
The first group of viruses had wholly Eurasian (EA) genome segments. The other three groups were reassortant viruses containing genome segments derived from both North American (NAm) and EA influenza A viruses.
Almost seventeen percent of the H5N1 viruses had mammalian adaptive mutations (E627K, E627V, and D701N) in the polymerase basic protein 2 (PB2) subunit of the RNA polymerase complex. Other mutations that may favour adaptation to mammalian hosts were also present in other internal gene segments.
The detection of these critical mutations in large number of mammals within short duration after virus introduction inevitably highlights the need for continually monitoring and assessing mammalian-origin H5N1 clade 2.3.4.4b viruses for adaptive mutations, which potentially can facilitate virus replication, horizontal transmission, and posing pandemic risks for humans.
(SNIP)
The current study describes the presence of wholly Eurasian and three novel reassortant H5N1 clade 2.3.4.4b viruses with various genome constellations in free-living mesocarnivore species (red foxes, striped skunks and American mink) in Canada in 2022. The viruses characterized from the mesocarnivore species had similar genetic compositions with H5N1 viruses circulating in avian species in the same geographic areas. The spillover of these viruses from wild birds to mammals could cause a potentially devastating pandemic if the H5N1 viruses mutate into forms that can spread efficiently among the mammalian species.
(SNIP)
Discussion
Influenza A viruses continuously evolve and as a result accumulate adaptive mutations, which facilitate virus replication, interspecies transmission, and pose pandemic risks for human and animals. In the current study, we examined forty HPAI H5N1 viruses from clade 2.3.4.4b virus with various genome constellations in multiple mammalian species, notably in mesocarnivore species in Canada.
Infections with HPAI H5N1 viruses and mortality in these free-living red foxes, striped skunks and mink reaffirm that these viruses can overcome interspecies barriers and infect new hosts. The clinical presentation of the disease in red foxes, skunks and mink was mostly neurologic, resembling clinical and pathologic features described in seals and foxes that were naturally infected with HPAI H5N8 viruses [25].
Oropharyngeal swabs and tissues consistently yielded positive PCR results for IAVs and fox kits that were recovered from the clinical infection developed anti-NP as well as anti HA-specific antibodies. Histologic lesions associated with HPAI H5N1 viruses in a number of red foxes include extensive meningoencephalitis and pneumonia and the presence of a higher load of viral antigen was demonstrated by IHC staining mainly in the brain sections. Reperant et al. (2008) experimentally infected red foxes with wild bird-origin HPAI clade 2.2 H5N1 virus via the intra-tracheal route and through ingestion [26]. Foxes inoculated by the intra-tracheal route excreted viruses orally for a relatively prolonged period of time and developed severe pneumonia, myocarditis and encephalitis.
Extensive amount of virus antigen was detected in the lung and brain of individuals infected intratracheally with the virus. The foxes that were infected by ingesting wild bird carcasses obtained from experimentally infected wild birds excreted the virus transiently and developed milder pneumonia with traces or undetected virus antigen in the lungs [26]. The observed differences in virus shedding, organ lesions and virus antigen load between the two modes of experimental infections may likely be related to the amounts of viruses in the inoculum and tissues of birds fed to the foxes.
Most infected mammals described in our study are opportunistic mesocarnivore species that were likely exposed to the H5N1 virus by consuming larger quantities of incapacitated or dead birds infected with H5N1 viruses. Fox kits from the same litter, likely shared the same infectious meal. However, horizontal transmission to den mates via aerosol or vertical transmission to kits due to consumption of milk from a lactating vixen can not be ruled out. A recent study in the UK provided a clue of possible transmission of HPAI H5N8 virus from swans to red foxes and seals by aerosol means while the birds and the mammalian species were kept in captivity in the same quarantine services [25].
While none of this tells us whether HPAI H5N1 can ever adapt well enough to spark a human pandemic, it does remind us that the virus is engaged in thousands of unregulated, and largely unseen gain-of-function experiments around the globe every day.
And persistence can be a mighty tool, even against long odds.