#17,688 Although we see sporadic cases every summer (and many more likely go undetected), spillovers of swine variant influenza viruses (typically H1N1v, H1N2v, H3N2v) are followed closely by health authorities because of their (currently low) potential to spark an epidemic or pandemic.
What is a variant influenza virus?
When an influenza virus that normally circulates in swine (but not people) is detected in a person, it is called a “variant influenza virus.” For example, if a swine origin influenza A H3N2 virus is detected in a person, that virus will be called an “H3N2 variant” virus or “H3N2v” virus.
Sporadic infections and even localized outbreaks among people with variant influenza viruses may occur. All influenza viruses have the capacity to change and it’s possible that variant viruses may change such that they infect people easily and spread easily from person-to-person. The Centers for Disease Control and Prevention (CDC) continues to monitor closely for variant influenza virus infections and will report cases of H3N2v and other variant influenza viruses weekly in FluView and on the case count tables on this website
Today, the CDC's FluView report contains the following description of a recent H1N2v case detected in Montana.
Novel Influenza A Virus
A human infection with a novel influenza A virus was reported by the Montana Department of Public Health and Human Services. The patient was infected with an influenza A(H1N2) variant (A(H1N2)v) virus. The patient was < 18 years of age at the time of infection, sought health care during the week ending August 5, 2023 (week 31), and has not been hospitalized. An investigation by local public health officials found that prior to their illness onset the patient attended an agricultural fair. The investigation is currently ongoing.
This is the third human infection with a variant influenza A virus reported in the United States in 2023, including one infection with an H3v (Michigan) virus and two infections with H1N2v (Michigan, Montana) viruses.
When an influenza virus that normally circulates in swine (but not people) is detected in a person, it is called a “variant” influenza virus. Most human infections with variant influenza viruses occur following exposure to swine, but human-to-human transmission can occur. It is important to note that in most cases, variant influenza viruses have not shown the ability to spread easily and sustainably from person to person.
Early identification and investigation of human infections with novel influenza A viruses are critical so that the risk of infection can be understood, and appropriate public health measures can be taken.
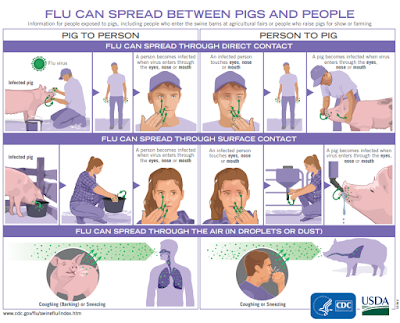
Additional information on influenza in swine, variant influenza virus infection in humans, and guidance to interact safely with swine can be found at www.cdc.gov/flu/swineflu/index.htm.
Additional information regarding human infections with novel influenza A viruses:
Surveillance Methods | FluView Interactive
Swine influenza surveillance is quite limited around the world, but the CDC's IRAT (Influenza Risk Assessment Tool) lists 3 North American swine viruses as having at least some pandemic potential (2 added in 2019).
H1N2 variant [A/California/62/2018] Jul 2019 5.8 5.7 Moderate
H3N2 variant [A/Ohio/13/2017] Jul 2019 6.6 5.8 Moderate
H3N2 variant [A/Indiana/08/2011] Dec 2012 6.0 4.5 Moderate
The CDC currently ranks a Chinese Swine-variant EA H1N1 `G4' as having the highest pandemic potential of any flu virus on their list.
Although it's encouraging that the 2009 H1N1 pandemic was the mildest in a century, there are no guarantees that a different novel H1N1 (or H3N2) virus would be as accommodating.