#15,788
Six years ago, in CDC: IRAT Evaluation Of Novel Avian & Swine Flu Risks, we looked at the CDC's Influenza Risk Assessment Tool which ranked the 11 most worrisome novel flu subtypes/strains that circulated in non-human hosts, and that posed a potential threat to human health.
Of particular note, 72% (8 of 11) had emerged since 2011, highlighting the rapid evolution and emergence of novel flu viruses over the past decade.
In 2017, 3 more viruses were added, and in june of 2018 the number rose to 16 (see CDC Adds Two New Novel Viruses To Their IRAT List). In May of 2020, three more viruses were added (2 swine-variant, 1 avian H9N2), bringing the total to 19.
The CDC uses two sets of criteria to evaluate novel viruses. One to estimate a virus's potential for sustained human-to-human transmission, and another to gauge it's potential for significant impact on public health.
At the top of the threats list for the past several years have been two avian H7N9 viruses which sparked major human outbreaks in China between 2013-2017. China's nationwide rollout of an H5+H7 poultry vaccine over the summer of 2017 has substantially reduced the risk of an outbreak.
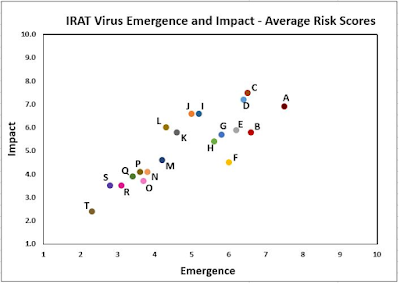
This week, however, the CDC updated their IRAT list, putting a 20th virus on the list - a swine variant EA H1N1 from Asia - which they rank with the highest (7.5) risk of emergence score, and the third highest impact score, we've seen to date.
H1N1: Eurasian avian-like Swine Influenza A(H1N1) [A/swine/Shandong/1207/2016] Virus
Human infections with Eurasian avian-like swine influenza A(H1N1) viruses (EA SIV H1N1) have been reported occasionally in China. Some infected persons reported direct or indirect exposure to swine. While first reported in Europe in 1979, EA SIV H1N1 reached China as early as 1993. Multiple genotypes have since been identified in China as a result of reassortment with other SIVs. A recent paper by Sun et al. described six genotypes (designated G1-G6) of EA SIV H1N1 detected in China between 2011 and 2018, with 3 human infections in which the G4 genotype was detected. The G4 EA SIV H1N1 virus has been reported to show efficient infectivity and transmissibility in the ferret model and a study in China showed that antibody titers against G4 viruses were higher in swine workers compared to the general population, suggesting that additional human infections may have occurred. Full genome sequences of the G4 viruses indicated the presence of EA SIV H1N1 HA and NA genes with a unique combination of internal genes including A(H1N1)pdm09 virus PB2, PB1, PA, NP and M genes and a triple reassortant NS gene.
Summary: A risk assessment of Eurasian avian-like swine influenza A(H1N1) [A/swine/Shandong/1207/2016] virus, clade 1C.2.3 and genotype 4, was conducted in July 2020. With point scores ranging from 1 to 10, the overall IRAT risk assessment score for this virus falls into the moderate risk category, which ranges from 4.0 to 7.9. The average risk score for potential emergence of the virus to achieve sustained human-to-human transmission was 7.5, within the upper moderate range. The average risk score for the virus to impact public health if it were to achieve sustained human-to-human transmission was 6.9, also in the upper moderate range. Full report here pdf icon[PDF – 272 KB].
We began following this particularly swine-influenza lineage back in late 2015, when Chen Hualan, director of China's National Avian Influenza Reference Laboratory and the lead author of a paper (see PNAS: The Pandemic Potential Of Eurasian Avian-like H1N1 (EAH1N1) Swine Influenza), publicly stated:
"Based on scientific analysis and comprehensive comparison of the main animal flu viruses: H1N1, H3N2, H5N1, H7N9, H9N2 and EAH1N1, we found the EAH1N1 is the one most likely to cause next human flu pandemic. We should attach great importance to the EAH1N1."
Researchers generated 55 novel reassortant viruses spread across 17 genotypes, demonstrating not only how readily EAH1N1 SIV can reassort with human H1N1pdm in a swine host, but also finding:
`Most of reassortant viruses were more pathogenic and contagious than the parental EA viruses in mice and guinea pigs'.
Later that same year, in EID Journal: Reassortant EAH1N1 Virus Infection In A Child - Hunan China, 2016, we reviewed the case report on a 30-month old child from Hunan Province, who was infected with one of these reassortant EAH1N1 - H1N1pdm viruses.
In 2017 we looked at J. Virology: A Single Amino Acid Change Alters Transmissibility Of EAH1N1 In Guinea Pigs, while in 2018 we saw Emerg. Microbes & Infect.: Effect Of D701N Substitution In PB2 Of EAH1N1 Swine Flu Viruses which described the growing diversity of novel novel H1N1 and H3N2 flu viruses (including EA H1N1) in China's pig population.
In 2019, we saw a number of studies on EAH1H1, including one that documented the virus in farmed mink in China (see Vet. MicroB.: Eurasian Avian-Like Swine Influenza A (H1N1) Virus from Mink in China).
A growing concern, last summer yet another study appeared in PNAS (see Eurasian Avian-like H1N1 Swine Influenza Virus With Pandemic Potential In China) that reported a greater than 10% seroprevalence for the EAH1N1 virus among swine workers tested, suggesting that EAH1N1 is gaining human infectivity.
This sparked a series of high profile risk assessments being published by the CDC, ECDC, WHO (and others) last summer.
The CDC's Responds to the PNAS EA H1N1 `G4' Swine Flu Study
ECDC Risk Assessment: Eurasian avian-like A(H1N1) swine influenza viruses
WHO Novel Flu Summary & Risk Assessment - July 2020
FAO/OIE/WHO Tripartite Statement on the Pandemic Risk of Swine Influenza
While the concern over the past two decades has been seeing the emergence of a high-mortality (10% CFR+) pandemic from avian H5Nx, H7Nx, or MERS-CoV, the lesson of COVID-19 is that even a 1%-2% CFR can set society back on its heels.
EA H1N1 has a good deal of competition, and so it isn't a slam dunk that this virus will spark the next influenza pandemic. Viruses can make abrupt strides when the evolutionary dice are tossed
But another influenza pandemic will surely come, perhaps sooner than we realize, and we need to be a lot better prepared for it than we were for COVID.